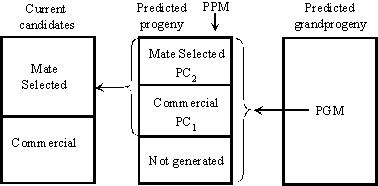
This is a paper by Ross and Brian which attempts to put mate selection into a useful framework for crossbreeding scenarios. It aims to generate both investment matings (to make eg. F1 females) as well as matings to capitalise on investments (to make eg. 3-breed crosses with F1 dams) in competition with each other.
Computational problems are such that Group Mate Selection will be needed for implementation.
A TACTICAL APPROACH TO
THE DESIGN OF CROSSBREEDING PROGRAMS.
R.K. Shepherd1 and B.P. Kinghorn2
1 Department of Mathematics and Computing, University of Central Queensland,
Rockhampton, Qld 4702, Australia
2 Department of Animal Science, University of New England, Armidale, NSW 2351, Australia
ABSTRACT
A Look Ahead Mate Selection (LAMS) scheme is developed with the objective of maximising phenotypic merit in each of the next two generations. This is of most value where non-additive effects are of importance, such as heterosis, inbreeding or non-additive QTL effects. The method is developed on the basis of a crossbreeding scenario, but it can be applied to any situation for which appropriate merit functions can be derived. Earlier work aimed just two generations ahead, but this can be suboptimal for non-additive effects, as ‘investment matings’ (such as generation of first cross females) are not properly exploited, due to lack of incentive to generate merit in the following generation. The method developed is tactical, in that it uses prevailing animal resources and generates a mate selection solution which constitutes the breeding decisions for the current mating. Integer programming and genetic algorithms can be used to find optimal solutions, but animal grouping strategies may be required to give realistic computing times.
INTRODUCTION
A tactical approach to making decisions about breeding for economic merit involves two steps:
The paper focuses on use of this approach when breed differences and heterosis are important, in addition to genetic variation within groups.
AIMING ONE OR TWO GENERATIONS AHEAD
If the objective is to maximise progeny genetic merit then mate selection can be carried out using Linear Programming (LP) techniques (Jansen and Wilton, 1985) or an approximate method (Kinghorn, 1986) which is less demanding computationally. Kinghorn and Shepherd (1994) extended the breeding horizon by aiming two generations ahead in order to maximise the genetic merit of the grandprogeny. The intention was to give due consideration to the exploitation of assortative mating and to enrich information in the pedigree structure by rearing relatives in different fixed effects groups, such as herd or management group ie. to breed for connection. An exchange algorithm was used to implement mate selection.
Aiming two generations ahead is very important for crossbreeding, most especially where maternal heterosis is important. For example, the optimum commercial animals two generations ahead may be the product of a three breed cross resulting from a terminal sire mated to an F1 female which is produced from matings in the current generation. Similarly a four breed cross can be aimed at two generations ahead by producing the two breed crosses from the current matings.
However, it may not be optimal to determine mate selection in each generation solely on the basis of maximising the predicted genetic merit in the grandprogeny. For example, consider a crossbreeding scenario without any within-breed additive genetic variation. If a three breed cross is optimum two generations ahead, the mating algorithm will set up F1 females in the next generation. However next generation when these animals are available for breeding, the mate selection algorithm may not select them because aiming two generations ahead the alternative mating to produce F1 females in the next generation may be optimal. The problem results from not distinguishing between matings to produce breeding animals (investment matings) and matings to produce commercial offspring (realisations of prior investments). This problem can be largely overcome by simultaneously considering merit both one and two generations ahead.

Figure 1.
AIMING BOTH ONE AND TWO GENERATIONS AHEAD
Figure 1 illustrates a Look Ahead Mate Selection (LAMS) scheme which carries out mate selection among predicted progeny, and, as a consequence mate selection among current candidates. Only commercial animals yield dollar returns from product each generation, and this is used to calculate predicted progeny merit (PPM) and predicted grandprogeny merit (PGM). The breeding objective is to maximise the combined return from commercial animals both one and two generations in the future. A full consideration would accommodate discounting of returns, which will not be done here, for simplicity. Some progeny (denoted by PC1) are predicted to be commercial animals in the next generation. Other progeny (denoted by PC2) are predicted to produce commercial animals two generations ahead, and are thus mate selected. In general, most predicted progeny are not generated, as the generation 0 mate selection set does not include their parents. Mate selection among predicted progeny also drives the mate selection of the current candidates, which is used to drive the breeding program.
Each generation, predicted progeny roles (PPR, which are either PC2 or PC1, or not generated) are determined by maximising the objective function. It should be emphasised that PC1 and PC2 are not sets of current generation mating pairs but rather sets of predicted progeny with predicted roles in later generations. A candidate pair for mating in the current generation may have some progeny predicted to be in PC1, and other progeny predicted to be in PC2. Each generation, PPR are determined according to predicted progeny merit (PPM), predicted grandprogeny merit (PGM), and the outcome of the optimisation algorithm, described below. This algorithm finds the mating set for the current generation which maximises the objective function, based on PPR and appropriate constraints according to the logistics of mating.
PREDICTED PROGENY AND GRANDPROGENY MERIT
To calculate PPM and PGM, we need genetic (and economic) models that
account appropriately for sources of variation. Consider a crossbreeding scenario in which
the only important sources of variation are within breed additive genetic variation (![]() ) and assumed known fixed between-breed effects: direct additive (
) and assumed known fixed between-breed effects: direct additive (![]() ), maternal additive (
), maternal additive (![]() ), direct dominance (
), direct dominance (![]() )
and maternal dominance (
)
and maternal dominance (![]() ) effects. The
predicted progeny merit (
) effects. The
predicted progeny merit (![]() ) is calculated as
follows:
) is calculated as
follows:
![]() (1)
(1)
![]() (2)
(2)
Using the same notation, the predicted grandprogeny merit (![]() ) is calculated as follows
) is calculated as follows
![]() (3)
(3)
where no within family variation is needed as no selection will be
applied. If epistatis is known to be important in explaining non-additive genetic
variation between breeds and good estimates exist then it should be included in PPM and
PGM. For example, additive x additive epistatic interaction would involve adding the term![]() to equations (1) and (2) where
to equations (1) and (2) where ![]() is the matrix of
between breed additive x additive epistatic effects - for example, E = (J - I)e assuming equality of breed epistatic effects
is the matrix of
between breed additive x additive epistatic effects - for example, E = (J - I)e assuming equality of breed epistatic effects ![]() , the maximum possible breakdown of favourable
purebreed epistatic effects.
, the maximum possible breakdown of favourable
purebreed epistatic effects.
MATE SELECTION ALGORITHM
Each pair of predicted progeny can have any of three roles: not
generated (ie. the parents are not mated), generated for PC1, or
generated and selected for PC2. Now the role value of the kth
predicted male progeny of the mating between candidates i and j and the rth
predicted female progeny of the mating between candidates p and q is ![]() for PC2 where G is the offspring of the kth
predicted male and rth predicted female, while for PC1 the
role value is
for PC2 where G is the offspring of the kth
predicted male and rth predicted female, while for PC1 the
role value is ![]() .
. ![]() (
(![]() ) if
the predicted progeny pair is selected for a PC2 (PC1) role,
otherwise
) if
the predicted progeny pair is selected for a PC2 (PC1) role,
otherwise ![]() (
(![]() ).
Deciding the role of a predicted progeny pair is not straightforward, as it depends on the
allocated roles of other predicted progeny.
).
Deciding the role of a predicted progeny pair is not straightforward, as it depends on the
allocated roles of other predicted progeny.
Let n be the number of progeny of each sex per dam and assume at most d dams per sire. If x mating pairs are required to be selected in each generation then the problem can be stated formally as follows, where PP is predicted progeny:
Maximise:
![]() subject to
subject to
No. of matings:
![]() (4)
(4)
![]() (5)
(5)
Individual PP:
![]() for each p, q, r (6)
for each p, q, r (6)
![]() for each i, j, k (7)
for each i, j, k (7)
![]() for each i, j, k (8)
for each i, j, k (8)
![]() for each i, j, k (9)
for each i, j, k (9)
Full sib PP:
![]() for each p, q (10)
for each p, q (10)
![]() for each i, j (11)
for each i, j (11)
Half sib PP:
![]() for each p (12)
for each p (12)
![]() for each i (13)
for each i (13)
![]() for each i (14)
for each i (14)
![]() for each i (15)
for each i (15)
The constraint equations can be interpreted as follows:
Eq. 4. There are exactly x PC2 matings. This constraint can be lifted, with the consequence that the emphasis on genetic merit in generations one and two will differ.
Eq. 5. There are exactly xn PP pairs selected for PC (PC1 and PC2) roles.
Eq. 6. Each female PP can be selected for at most one role.
Eq. 7. Each male PP can be allocated at most d mates for PC2.
Eq. 8. Each male PP can be selected for at most one PC1 role.
Eq. 9. Each male PP can be selected for a role as either PC1 or PC2 but not both.
Eq. 10. Full sib female PP either are not generated (= 0) or are selected for PC roles (= n).
Eq. 11. Full sib male PP either are not generated (= 0) or are selected for at least n PC roles.
Eq. 12. For male candidate p there are at most d matings, each generating n female PP.
Eq. 13. For male candidate i there are at most d matings, each generating n male PP who could be allocated PC1 roles.
Eq. 14. For male candidate i there are at most d matings, each generating n male PP, with each able to be used in at most d PC2 roles.
Eq. 15. For male candidate i there are at most d half sib families with all n female PP selected for PC roles
To put some emphasis on controlling inbreeding, further constraints
could be added. For example, to not allow any predicted full sib progeny to be mated, add
the constraint ![]() for each i, j combination.
Alternatively, a proper modelling of inbreeding depression in PPM and PGM will give mate
selection to reduce inbreeding coefficients in the next two generations.
for each i, j combination.
Alternatively, a proper modelling of inbreeding depression in PPM and PGM will give mate
selection to reduce inbreeding coefficients in the next two generations.
An LP could solve this problem, except for constraints (9), (10) and
(11). Constraint (9) would vanish for single pair mating (![]() ), if aiming only one generation ahead (only PC1) or if
aiming only two generations ahead (only PC2). The (or
), if aiming only one generation ahead (only PC1) or if
aiming only two generations ahead (only PC2). The (or ![]() ) option in constraints (10) and (11) could be
removed by only considering PC role allocation, rather than including PP which are not
generated. That is, given a particular parental mating set, find the best
allocation of roles to predicted progeny in order to maximise the objective.
) option in constraints (10) and (11) could be
removed by only considering PC role allocation, rather than including PP which are not
generated. That is, given a particular parental mating set, find the best
allocation of roles to predicted progeny in order to maximise the objective.
However, even as an LP, the problem is still too large. For example,
with 1000 (m) males and 1000 (f) females available for selection and 1 (n)
progeny of each sex per dam then the total number of possible predicted progeny is ![]() (mfn) and there would be
(mfn) and there would be ![]() (2 x mfn x mfn) unknown
binary variables in the matrices X and Y. The algorithm of Kinghorn (1986)
would have great computational efficiency here. Hence in general the task is an integer
nonlinear programming (NLP) problem which is not computationally feasible for typical
crossbreeding schemes. Good approximate methods are required to solve this mate selection
problem.
(2 x mfn x mfn) unknown
binary variables in the matrices X and Y. The algorithm of Kinghorn (1986)
would have great computational efficiency here. Hence in general the task is an integer
nonlinear programming (NLP) problem which is not computationally feasible for typical
crossbreeding schemes. Good approximate methods are required to solve this mate selection
problem.
EVOLUTIONARY ALGORITHMS AND GROUP MATE SELECTION
Genetic Algorithms, GA’s, and the more recent Evolutionary
Algorithms, EA’s (Michalewicz, 1994) mimic biological evolution in their quest to
find the best solution for a complex optimisation problem. The earlier NLP implicitly
selects between all possible matings sets which is usually a very large number (see box
below). However many of these mating sets are not very competitive for the objective.
Hence an alternative approach would be to use an EA to search among competitive mating
sets in generation 0. Then for each mating set evaluated, the NLP problem becomes one of
mate selection in generation 1. For a particular generation 0 mating set the number of
predicted progeny reduces to ![]() per sex and the
number of elements in the matrices X and Y reduces to
per sex and the
number of elements in the matrices X and Y reduces to ![]() , a much smaller NLP problem. For example, for m = f = 1000,
n = 1 and x = 1000 the number of elements reduces to
, a much smaller NLP problem. For example, for m = f = 1000,
n = 1 and x = 1000 the number of elements reduces to ![]() . An EA could be designed to solve the mate selection problem
directly. However it is probably more time efficient to use an EA to search among possible
mating sets in conjunction with a different EA designed to solve the generation 1 mate
selection problem for each mating set. Hayes et al (1998) discuss the application of
genetic and evolutionary algorithms to the mate selection problem.
. An EA could be designed to solve the mate selection problem
directly. However it is probably more time efficient to use an EA to search among possible
mating sets in conjunction with a different EA designed to solve the generation 1 mate
selection problem for each mating set. Hayes et al (1998) discuss the application of
genetic and evolutionary algorithms to the mate selection problem.
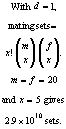
One approach to reducing computation time is to group animals and to run a mate selection algorithm at the level of groups, leading to a solution describing the number of animals to be selected out of each group and the pattern of mate allocation (Kinghorn, 1998). This can be followed by a ‘fine tuning’ individual animal mate selection using only animals selected out of groups. This 2-step approach gave high efficiency yet dramatically lower computing times (Kinghorn, 1998).
EXAMPLE
The example relates to a tiny problem for which there is one male and one female candidate of each of breeds 1 and 2 in generation 0. Two predicted progeny of each sex (n = 2) are generated for each of the four candidate matings – eg. males 11a and 11b, and females 11c and 11d from the mating 1 x 1 (male x female). Figure 2 illustrates all possible matings among these predicted progeny.
Table 1 shows the PPM for predicted progeny and PGM for their crosses. For simplicity of presentation in this table, matrix l is replaced by a column of PPM values for males and a row of PPM values for females. This leads to an alternative set of constraints, sufficient for this example and described in the caption of Table 1. For two single pair matings to be selected in each of generations 0 and 1 (x = 2, d = 1), one optimal solution is matings 1 x 2 and 2 x 1 in generation 0, matings 12b x 21d and 21b x 12d in generation 1, and PC1 roles for progeny 12a and 12c.
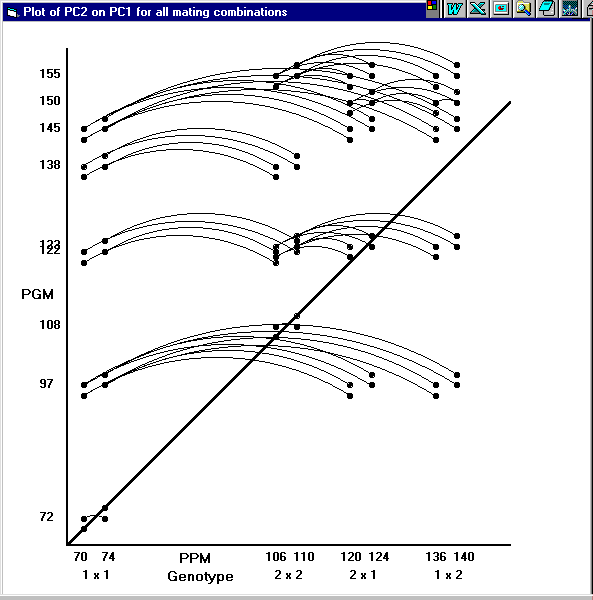
Figure 2. Plot of PGM (predicted grandprogeny merit) on PPM (predicted progeny merit) for the example. All possible individual matings between predicted progeny are shown. Groups of four points at the diagonal represent matings within purebreeds (and within full sib families in this tiny example). Matings exactly on the diagonal are between predicted progeny of equal breeding value, and those just off the diagonal are between predicted progeny of unequal breeding value. All matings between predicted progeny of different PPM are linked by an arc. PGM are generally higher than PPM because, in this example, maternal heterosis is only expressed in generation 2.
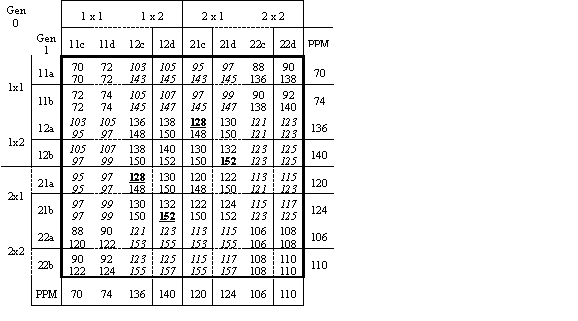
Table 1. Average PPM (top within cell) and PGM (bottom within cell) both within the bold frame, for the example. Male (female) predicted progeny are listed in the second column (row). PPM values are also shown in the right (male) and bottom (female) margins. Ad1=80, Ad2=100, Am1=-8, Am2=8, Dd=40, Dm=40. Predicted progeny a and c (b and d) have breeding values of +2 (-2) as deviations from breed and family means. Given d=1, constraints on solutions (0 or 1 for each location within each cell) are: each generation 0 row plus matched column S =0 or S =2n, each generation 1 row or column S £ 1, and (optionally, see comment on eq.4) total PC2 S =x. Cells in italics are not feasible under single-pair mating. A solution for x= 2 matings per generation is shown in bold and underlined.
DISCUSSION
As presented in this paper, each predicted progeny pair is chosen on the basis of either PPM or PGM - analogous to use of independent culling levels (ICL). This will not be generally optimal. For example, consider two PP pairs, both with PGM > PPM, with one pair marginally superior in PGM but markedly inferior in PPM. The pair with marginally lower PGM should be selected, given the potential prospect of PC1 roles for these animals. This is especially true if the accuracy of genetic parameters to estimate PPM and PGM is not high. However, using a simple index of PPM and PGM as the criterion of merit, rather than ICL, will miss the power that this approach has for setting up matings specifically targeted either one or two generations ahead. One possible solution is to use a composite index which is a weighted average of ICL and a simple index. A related issue arises when product can be harvested from breeding animals, in which case PC2 matings contribute product in the next generation.
With proper specification and modelling of the objective function, the method presented here has prospects of being able to exploit non-additive QTL effects, exploit assortative mating, avoid inbreeding depression one and two generations ahead, and set up a 4-pathway progeny testing scheme and various MOET schemes (where flexibility to choose fecundity is modelled).
The present method aims to maximise the sum of merit in commercial offspring in generations 1 and 2 alone. It ignores later generations, which is a weakness. For example, increasing the breeding values of a pair of predicted progeny has an equal impact on their average PPM and on the PGM if they were to be mated. This means that the ‘breeding line’ has no more power to attract high EBV animals than the ‘commercial line’. This could be overcome by extending the objective function to give appropriate weighting to long-term genetic merit, and other factors such as inbreeding level. Relative weightings over time will differ between factors. For example, there might be short-term emphasis on expression of heterosis and favourable QTL genotypes, but longer term emphasis on polygenic breeding value and optimal QTL allele frequencies.
ACKNOWLEDGEMENTS
Much of this work was conducted during a visit by the authors to the Roslin Institute and the Scottish Agricultural College, Edinburgh, with funds provided by the Australian DIST and the UK BBSRC Underwood Fund.
REFERENCES
Jansen, G.B. and Wilton, J.W. (1985) J. Dairy Sc. 68:1302-1305
Goddard, M.E. and Howarth, J.M. (1995) Proc. 5th WCGALP 18:306-309
Hayes, B.J., Shepherd, R.K., Newman, S. and Kinghorn, B.P. (1998) Proc. 6th WCGALP
Kinghorn, B.P. (1986) Proc. 3rd WCGALP 12:233-244
Kinghorn, B.P. (1998) J. Dairy. Science. Submitted.
Kinghorn, B.P. and Shepherd, R.K. (1990) Proc. 4th WCGALP 15:7-15
Kinghorn, B.P. and Shepherd, R.K. (1994) Proc. 5th WCGALP 18:255-261
Michalewicz, Z (1994) Genetic algorithms + data structures = evolution programs 2nd edition, Springer-Verlag, Berlin.